Challenges in Deep Learning-Based Small Organ Segmentation: A Benchmarking Perspective for Medical Research with Limited Datasets
Headline result
On only nine annotated histology images, foundation models retain performance under distribution shift while classical architectures collapse. Bootstrap confidence intervals overlap so substantially among top models that ranking differences are mostly statistical noise rather than algorithmic superiority.
Method in brief
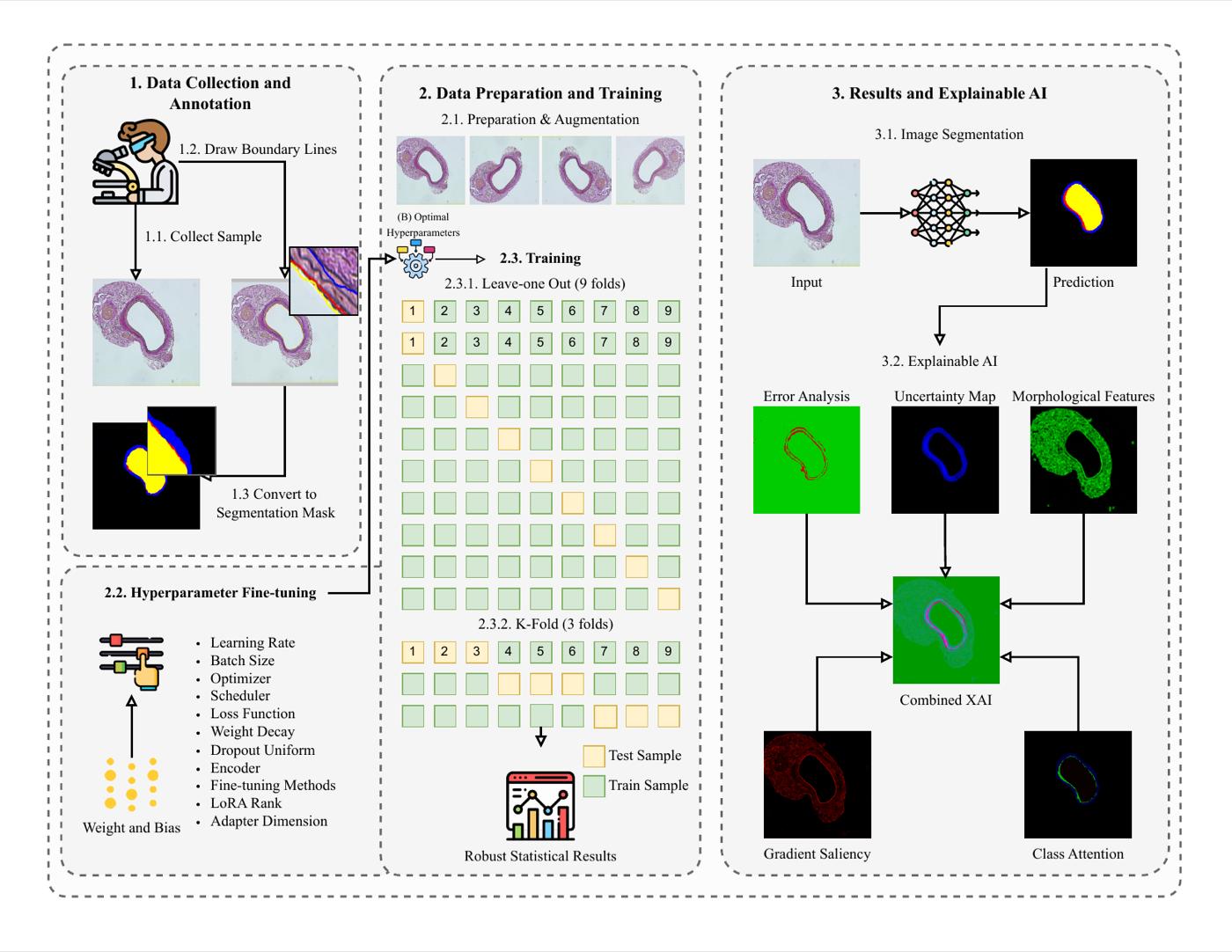
Ten segmentation models (classical, modern CNN, Vision Transformer, foundation models) are evaluated on a limited dataset of nine cardiovascular histology images, with ablations on data augmentation, input resolution, and seed stability. Generalisation is tested on an independent N = 153 set under distribution shift, with bootstrap confidence intervals quantifying the variance from algorithmic vs. statistical sources.
Key Contributions
- Systematically evaluates ten segmentation models (classical, modern CNNs, Vision Transformer, foundation models) on only nine annotated cardiovascular histology images.
- Ablation studies on data augmentation, input resolution, and random seed stability quantify the sources of variance in low-data benchmarking.
- Independent generalisation set (N = 153) under distribution shift exposes that rankings change substantially between in- and out-of-distribution evaluation, with foundation models retaining performance where classical architectures collapse.
- Bootstrap confidence intervals overlap substantially among top models, demonstrating that apparent ranking differences are largely statistical noise — and motivating uncertainty-aware evaluation as standard practice for low-data clinical research.
Abstract
Accurate segmentation of carotid artery structures in histopathological images is vital for cardiovascular disease research. This study systematically evaluates ten deep learning segmentation models including classical architectures, modern CNNs, a Vision Transformer, and foundation models, on a limited dataset of nine cardiovascular histology images. We conducted ablation studies on data augmentation, input resolution, and random seed stability to quantify sources of variance. Evaluation on an independent generalisation dataset (N = 153) under distribution shift reveals that foundation models maintain performance while classical architectures fail, and that rankings change substantially between in-distribution and out-of-distribution settings. Training on the second dataset at varying sample sizes reveals dataset-specific ranking hierarchies, confirming that model rankings are not generalisable across datasets. Despite rigorous Bayesian hyperparameter optimisation, model performance remains highly sensitive to data splits. The bootstrap analysis reveals substantially overlapping confidence intervals among top models, with differences driven more by statistical noise than algorithmic superiority. This instability exposes limitations of standard benchmarking in low-data clinical settings and challenges assumptions that performance rankings reflect clinical utility. We advocate for uncertainty-aware evaluation in low-data clinical research scenarios from two perspectives: first, the scenario is not niche and is rather widely spread; and second, it enables pursuing or discontinuing research tracks with limited datasets from incipient stages of observations.